A genetic screen identifies mutations in the yeast WAR1
gene, linking transcription factor phosphorylation
to weak-acid stress adaptation
Christa Gregori*, Bettina Bauer*, Chantal Schwartz, Angelika Krenà, Christoph Schu
¨ller§
and Karl Kuchler
Medical University Vienna, Max F. Perutz Laboratories, Department of Medical Biochemistry, Campus Vienna Biocenter, Austria
Weak acids have a long history as additives in food
preservation. In addition to sulfites used in wine mak-
ing, acetic, sorbic, benzoic and propionic acids are
commonly used in the food and beverage industry
to prevent spoilage [1,2]. In solution, weak acids exist
in a dynamic equilibrium between undissociated,
uncharged molecules and their anionic form. These
acids display increased antimicrobial action at low pH,
which favors the undissociated state. The uncharged
molecules can readily diffuse through the plasma
Keywords
ABC transporter; stress response; weak
organic acids; yeast; zinc finger
Correspondence
K. Kuchler, Medical University Vienna, Max
F. Perutz Laboratories, Department of
Medical Biochemistry, Campus Vienna
Biocenter, Dr Bohr-Gasse 9 ⁄2, A-1030,
Vienna, Austria
Fax: +43 1 4277 9618
Tel: +43 1 4277 61807
E-mail: karl.kuchler@meduniwien.ac.at
Present address
Universite
´de Nice-Sophia Antipolis,
Inserm, U636, Centre de Biochimie, UFR
Sciences, Parc Valrose, Nice, France
àInstitute of Biochemistry and Genetics,
Department of Clinical and Biological
Research (DKBW), Basel, Switzerland
§University of Vienna, Max F. Perutz
Laboratories, Department of Biochemistry &
Molecular and Cellular Biology, Campus
Vienna Biocenter, Austria
*
These authors contributed equally to this
work
(Received 11 January 2007, revised 4 April
2007, accepted 19 April 2007)
doi:10.1111/j.1742-4658.2007.05837.x
Exposure of the yeast Saccharomyces cerevisiae to weak organic acids such
as the food preservatives sorbate, benzoate and propionate leads to the
pronounced induction of the plasma membrane ATP-binding cassette
(ABC) transporter, Pdr12p. This protein mediates efflux of weak acid ani-
ons, which is essential for stress adaptation. Recently, we identified War1p
as the dedicated transcriptional regulator required for PDR12 stress
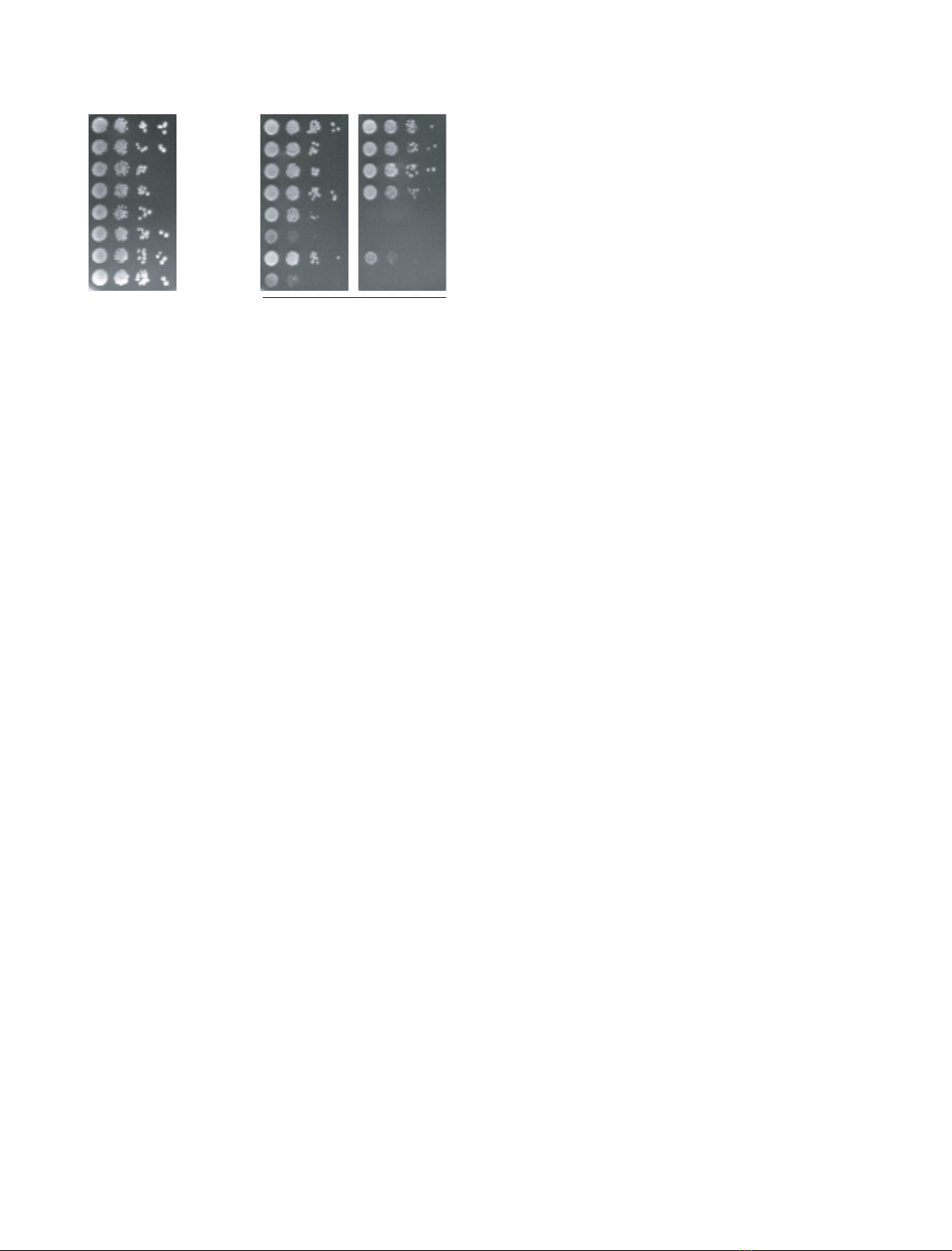
induction. Here, we report the results from a genetic screen that led to the
isolation of two war1 alleles encoding mutant variants, War1-28p and
War1-42p, which are unable to support cell growth in the presence of sorb-
ate. DNA sequencing revealed that War1-28 encodes a truncated form of
the transcriptional regulator, and War1-42 carries three clustered mutations
near the C-terminal activation domain. Although War1-42 is expressed and
properly localized in the nucleus, the War1-42p variant fails to bind the
weak-acid-response elements in the PDR12 promoter, as shown by in vivo
footprinting. Importantly, in contrast with wild-type War1p, War1-42pis
also no longer phosphorylated upon weak-acid challenge, demonstrating
that phosphorylation of War1p, its activation and DNA binding are tightly
linked processes that are essential for adaptation to weak-acid stress.
Abbreviations
GST, glutathione S-transferase; MHR, middle homology region; NLS, nuclear localization signal; PDR, pleiotropic drug resistance; WARE,
weak-acid-response element; YPD, yeast peptone ⁄dextrose.
3094 FEBS Journal 274 (2007) 3094–3107 ª2007 The Authors Journal compilation ª2007 FEBS